Research Interests
I. Computational Structural Biophysics and Systems Biology. My current and also previous research projects are broadly in the area of structural biophysics and systems biology. Presently, at NYU, in my role as a senior research scientist, I primarily support the research projects of various labs which spans from systems/genomics data analysis to structural biophysics, simulations and predictions.
In my previous position at Columbia University, I was mainly involved in few projects related to the development of computational methods to analyze and predict the RNA-protein binding interactions, RNA-protein complex structures using crosslinking, sequence and structural datasets.
II. Computational Structural Biophysics. My primary research area has been computational structural biology and biophysics. In my previous position in Dr. Joachim Frank's lab (04/2015 - 08/2020), I was mainly involved in the development of methods to study the structure and dynamics of macromolecular machines using cryo-EM image datasets. Following are the details of the individual projects:
1. Continuum of conformational states from cryo-EM data using manifold learning:
(a) I was working on a method ManifoldEM to obtain a continuum of conformational states of macromolecular machines from cryo-EM data.
Conventional methods using maximum likelihood classification, analyzes the projection images and determines the 3D structures with various conformation.
Hoewever, these methods are able to determine only a limited number of discrete classes.
Our approach aims to determine the continuum of conformational states (landscape) given enough projection images
of the molecular machines (Dashti et al. 2014, Frank et al. 2016). We achieve this task by analyzing the "space of images" employing the methodology known as manifold learning.
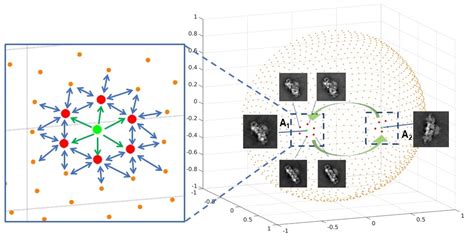
(b) Propagation of reaction coordinates across angular space:One important step in deriving a consolidated map of occupancies and energy landscape, is to propagate the reaction coordinates, from a known "anchor PD" and this has been done thus far "by hand".
In this work we have developed a method (Accepted to Journal of Chemical Information and Modeling, Special Issue:Frontiers in CryoEM Modeling, Maji et al. 2020), that attempts to solve the propagation problem by using a probabilistic graphical model and feature extraction methods such as optical flow & histogram of oriented gradients.
We illustrate the successful application of the propagation method to a ryanodine receptor and ribosome data.
2. Structure and function of 30S translational initiation complex: Recently, I got involved in an ongoing project in our group using cryo-EM to obtain insight about the conformational rearrangements of the ribosomal subunits during the initiation process.
3. Characterization of domain motion with coordinate transformation using moments of inertia tensor: I am also working on developing methods to study the domain motion of macromolecular structures. Molecular machines such as the ribosome,
while performing a certain biophysical process (e.g. translation, protein synthesis , etc.), undergo a series of conformational changes often characterized by relative motions of their domains.
These processes can be studied using various structural biology techniques and usually we have conformational snapshots of the molecular machines (pdb structures), obtained from flexible fitting of low-resolution cryo-EM density maps,
from X-ray crystallography, or from molecular dynamics simulation trajectories. The goal is to analyze these snapshots and obtain meaningful information about the motions of the molecular machines.
We have recently developed a comprehensive method (Maji et al. 2017) implemented
on the VMD platform, which can quantify these motions in a general and biologically informative way. Our objective is to characterize the motion of each domain as a
coordinate transformation using moments of inertia tensor (Agirrezabala et al. 2011, 2012). These tools presents a practical way for reproducible segmentation of domains,
and provide a generalized description of their motions
using principal axes of inertia. We can obtain the rotation angle and the native rotation axis for certain domain undergoing motion. Moreover, we can also determine
the component of the rotation around a specified axis, not necessarily the native rotation axis. Using such a toolset we can compare domain motion data from different researchers.
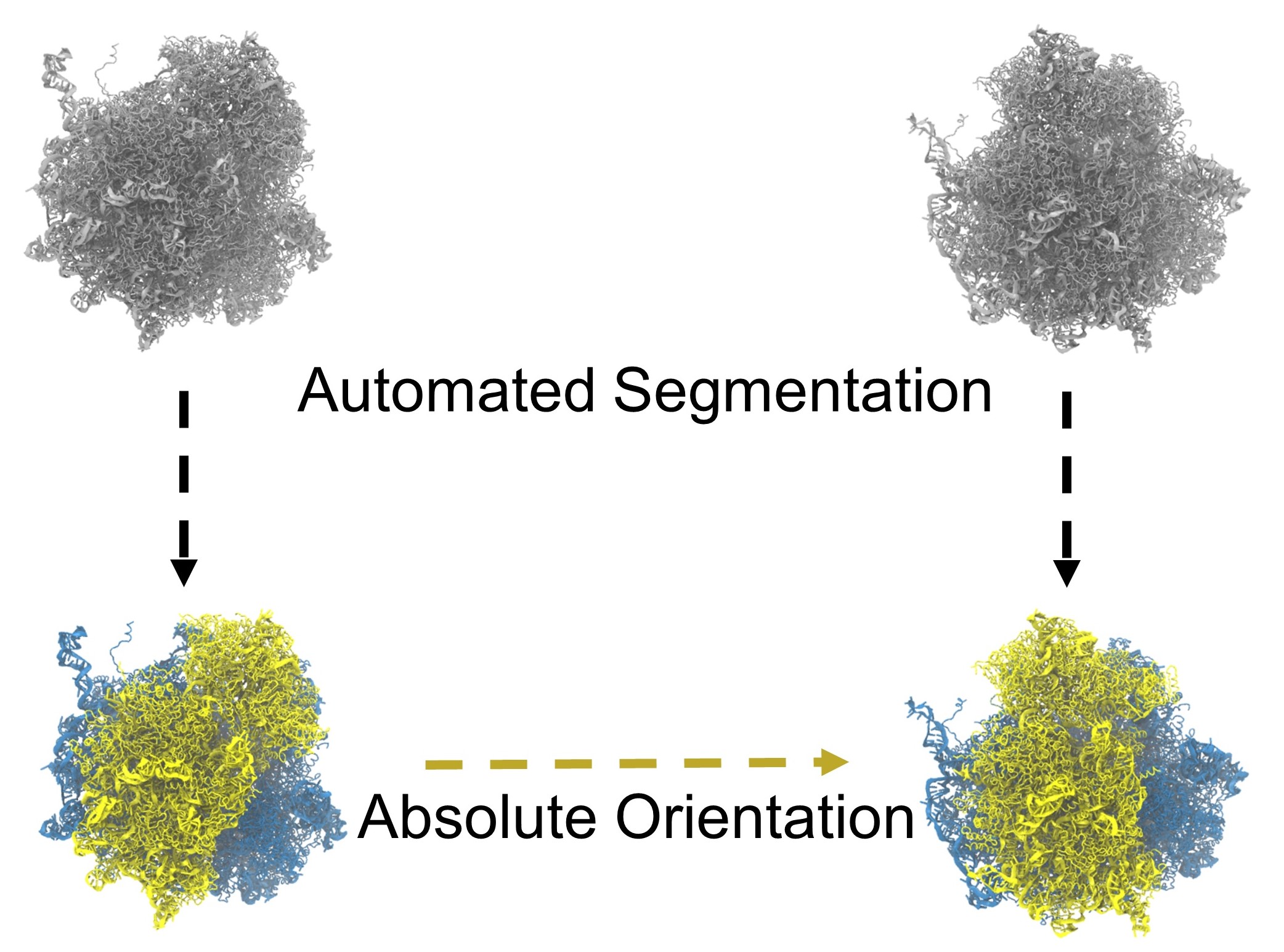
Quantifying the domain motion of macromolecular machines

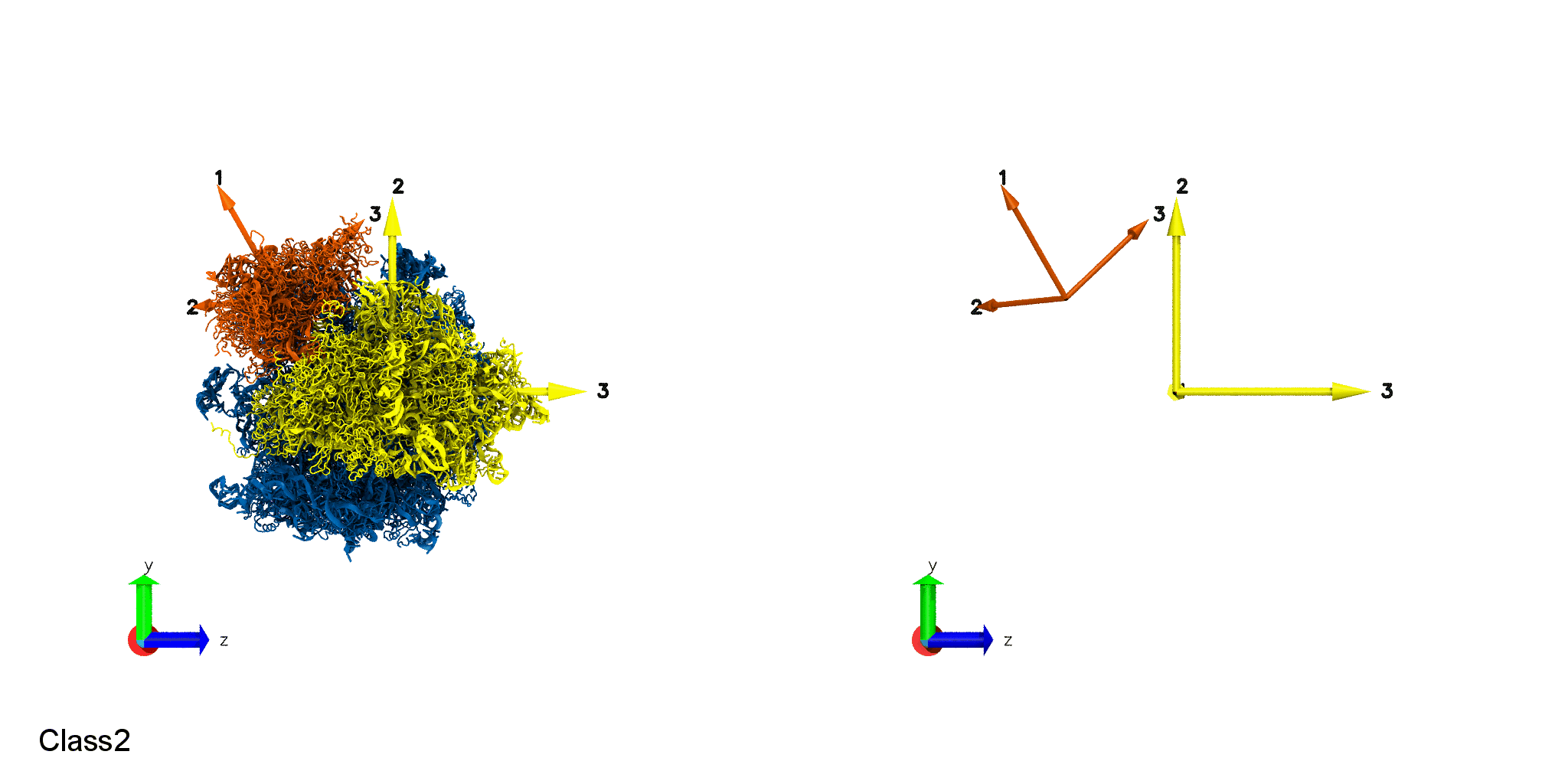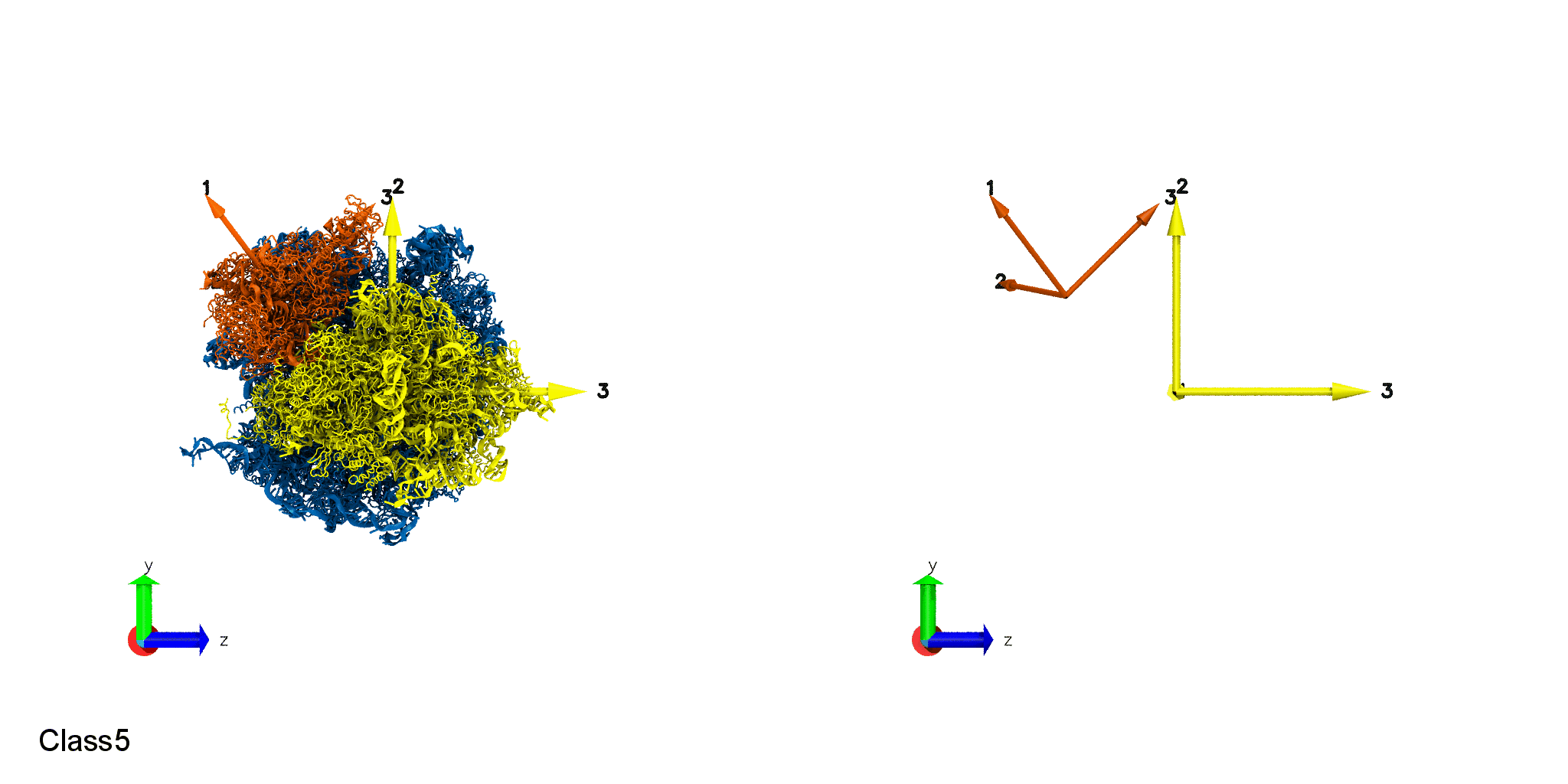
Domain Motion of a molecular machine, ribosome. Small Subunit Body rotation (Top). Small Subunit Head rotation (Bottom).
Reference: Quantitative Characterization of Domain Motions in Molecular Machines
DOI: 10.1021/acs.jpcb.6b10732
Previous Research
My previous research interests were single molecule Biophysics and fluorescence microscopy image analysis. I was involved in few different projects such as single particle tracking of biological molecules and superresolution bioimaging studies.
My main project was on developing approaches using Generative models for inferring biological structures from single molecule super-resolution datasets, which can potentially improve dynamic localization imaging of structural components in cells.
I was also involved in the development of three-dimensional single molecule localization algorithms using fluorescent beads, quantum dots and fluorogen activating peptides (FAPs).
Apart from single molecule imaging projects, I have also worked on understanding co-translational folding of protein, where I computationally studied the effect of codon usage and tRNA concentration on the translation rates for several proteins.
Research Summary
IV. General Topics